Oncology relational dataset derived from MIMIC-IV
This project presents a curated, oncology-focused relational dataset derived from the MIMIC-IV v3.1 clinical database, one of the largest publicly available collections of longitudinal, patient-level hospital data from the Beth Israel Deaconess Medical Center.
While MIMIC-IV is not an oncology cohort and contains a limited number of cancer patients relative to dedicated registries, it provides a rich clinical environment for exploring how data can be structured, queried, and integrated at scale.
The dataset isolates and organises clinical signals relevant to cancer patients to support exploratory analysis, visualisation, and methodological machine learning workflows. It is designed as a technical foundation for bridging routine clinical data (MIMIC-IV) with external molecular resources (e.g. TCGA).
Workflow
Raw Clinical Tables → Oncology Filtering → Signal Selection → Derived Oncology Cohort → Dataset
-
Extract core clinical tables
Imported key MIMIC-IV tables (patients,admissions,diagnoses_icd,icustays,prescriptions,labevents) into a local PostgreSQL environment -
Identify oncology-relevant records
Filtered diagnoses using cancer related ICD-9 (140–239) and ICD-10 (C00–D49) codes to define the clinical scope of the dataset -
Select relevant laboratory and treatment signals
Retained laboratory variables such as blood counts, liver enzymes, and selected tumour markers, together with medications linked to chemotherapy, immunotherapy, or hormonal treatment -
Define a derived oncology cohort
Built an ICD-based cohort table to enable efficient filtering and reproducible patient-level selection across downstream analyses -
Create supporting layers
Added derived views for treatment timelines and labelled laboratory values to facilitate longitudinal interpretation and exploratory modelling -
Optimise for querying and reuse
Applied indexes and materialised views to support interactive querying, dashboarding, and future analytical extensions
Metadata
Metadata
- Source: MIMIC-IV v3.1
- Institution: Beth Israel Deaconess Medical Center (2008–2022)
- Modules Used:
hosp,icu - Programming: PostgreSQL, Python (ETL), SQL (views and indexing)
- Focus: Oncology-relevant EHR subset
- Primary Cohort Definition: Patients with ICD-9/10 neoplasm codes
- Compliance: PhysioNet DUA 1.5.0, HIPAA-deidentified, time-shifted
Core Schema Structure
Schema Structure
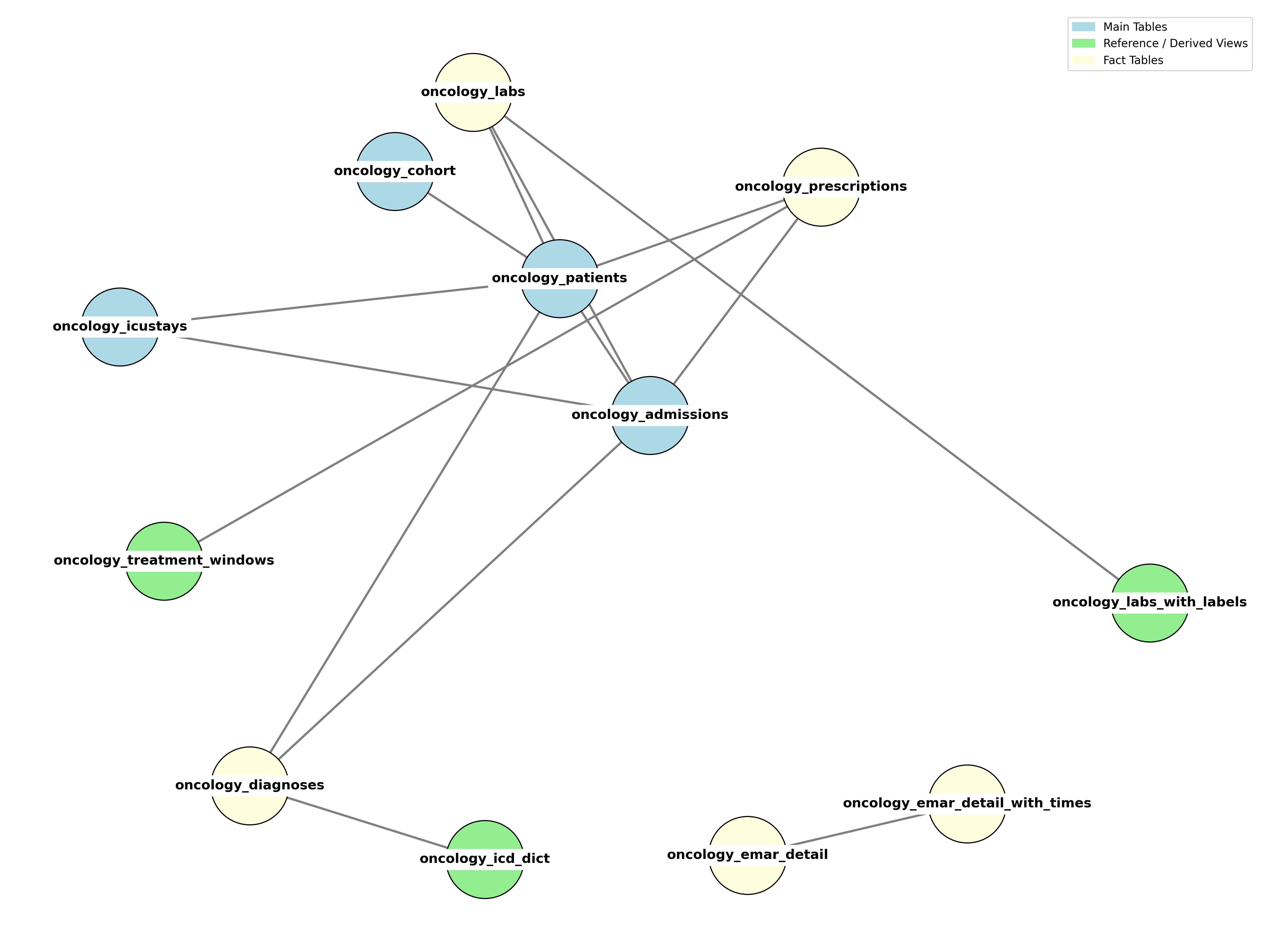
The dataset is organised into three complementary table categories:
| Category | Description | Examples |
|---|---|---|
| Main Tables | Core entities: patients, admissions, ICU stays, cohort flags | oncology_patients, oncology_admissions, oncology_icustays, oncology_cohort |
| Fact Tables | Event-level clinical records: diagnoses, labs, prescriptions, medication admin | oncology_diagnoses, oncology_labs, oncology_prescriptions, oncology_emar_detail |
| Reference Tables | Dictionaries and derived views for enrichment and summarisation | oncology_icd_dict, oncology_labs_with_labels, oncology_emar_detail_with_times, oncology_treatment_windows |
Use Cases
- Track cancer patients longitudinally (laboratory values, medications, ICU exposure)
- Prototype outcome or response modelling pipelines (methodological scope)
- Build dashboards and exploratory clinical analytics
- Practice reproducible clinical data engineering and SQL workflows
- Integrate clinical trajectories with external genomics resources (e.g. TCGA)