Translational Knowledge Graph for Oncology Target Prioritization
This project presents a translational knowledge graph designed to support early-stage oncology target assessment by integrating pharmacological and transcriptomic evidence within a single semantic framework.
As a proof of concept, it connects:
- Kinase bioactivity data from ChEMBL
- Normal tissue expression from the GTEx project
- Tumor expression from The Cancer Genome Atlas (TCGA)
- Identifier harmonisation via UniProt mappings to Ensembl genes and the UBERON anatomy ontology
Integration Overview
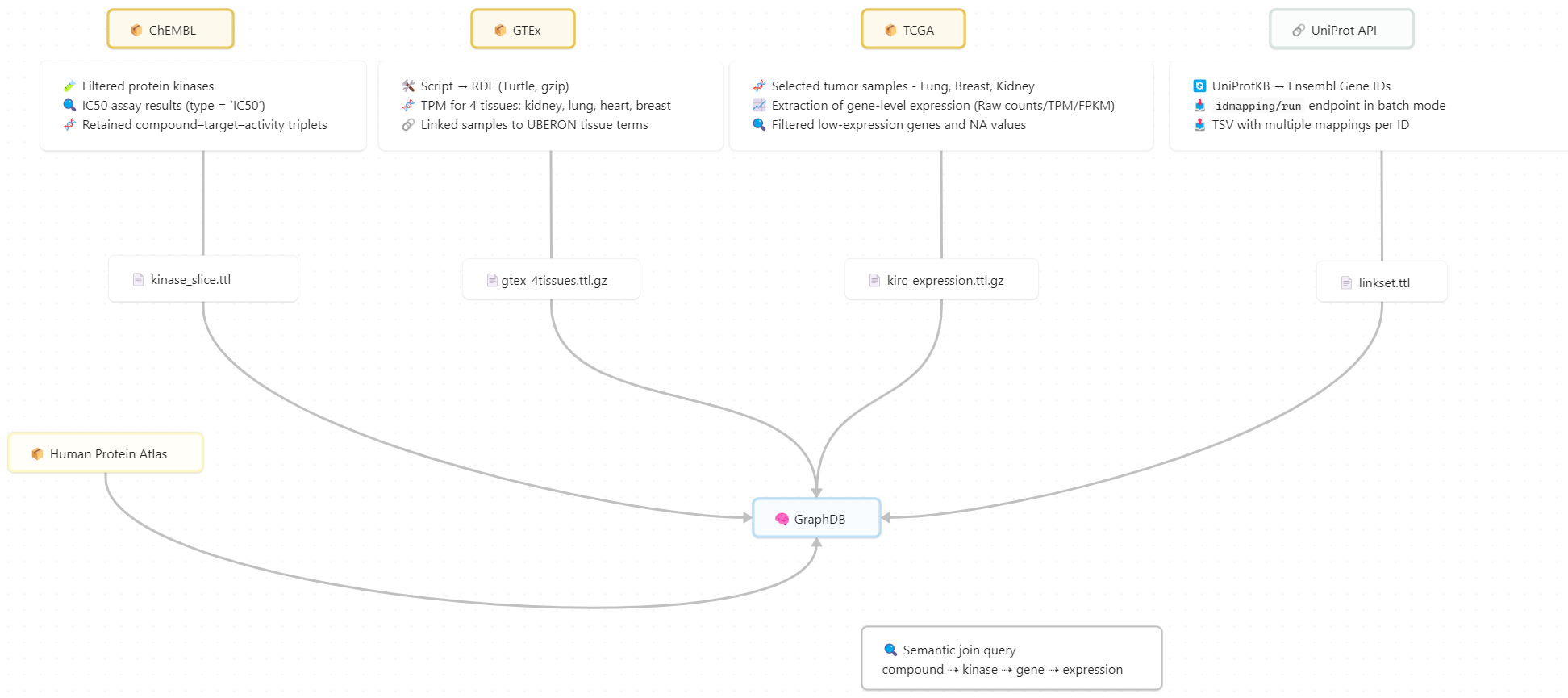
Each source dataset is processed independently, exported to Turtle (TTL), and loaded into a GraphDB triple store as separate named graphs.
Cross-dataset relationships are harmonised through shared identifiers, including UniProt ↔ Ensembl mappings and anatomical terms such as UBERON, enabling explicit semantic joins across pharmacological and transcriptomic datasets.
Because the integrated data are represented as RDF, they can be queried using SPARQL, the standard query language for semantic graphs.
What the Graph Enables
This structure supports biologically meaningful questions across compounds, genes, and normal versus tumor expression profiles, such as:
- "Which compounds inhibit proteins whose genes are overexpressed in tumors but not in healthy tissues?"
- "Which genes show tumor-specific expression patterns across tissues that have been previously targeted?"
Example use cases include:
- Prioritising targets based on tumor-specific expression patterns
- Derisking candidate targets by filtering for expression in healthy tissues as a toxicity proxy
- Ranking compounds according to the expression of their target set in a selected cancer type
- Comparing tumour versus normal tissue expression across tissues to assess specificity
- Assessing compound–target coverage to distinguish characterized proteins from more novel candidates
The Human Protein Atlas is a planned extension to incorporate information at protein level.